[1]:
# the boring stuff:
import numpy
import pandas
import scipy
from matplotlib import pyplot
from IPython.display import display, Image
# probabilistic programming:
import arviz
import pymc
import sunode
import pytensor
import pytensor.tensor as pt
import pytensor.printing
# modeling:
import calibr8
import murefi
Differentiable ODE models with murefi and sunode
This example builds on top of the basic Michaelis-Menten example to showcase how a murefi ODE model can be auto-differentiated with sunode. As we will see at the end of this example, this enables us to run efficient MCMCs with NUTS.
Preparing Data & Models
The next cell is a condensed version of the corresponding chapter from the basic example (Example_03_MichaelisMenten.ipynb).
[2]:
class ProductAssayModel(calibr8.BasePolynomialModelT):
def __init__(self):
super().__init__(
independent_key='P',
dependent_key='A570',
mu_degree=1,
scale_degree=0,
)
def observe_with_true_parameters(x):
return scipy.stats.t.rvs(loc=0.17 + 0.12 * x, scale=0.03, df=3)
# generate fake calibration data with a linear relationship
numpy.random.seed(202103)
N = 48
X = numpy.linspace(0.1, 10, N)
Y = observe_with_true_parameters(X)
cm_product = ProductAssayModel()
theta_fit, _ = calibr8.fit_scipy(
cm_product,
independent=X, dependent=Y,
theta_guess=[0, 0.2, 0.05, 5],
theta_bounds=[(-2, 2), (0.001, 0.5), (0.001, 0.5), (1, 10)]
)
fig, axs = calibr8.plot_model(cm_product);
for ax in axs:
ax.set_xlabel('product concentration [mmol/L]')
axs[0].set_ylabel('$A_{570}$ [a.u.]')
axs[1].set_ylabel("")
axs[0].set_ylim(bottom=0)
axs[1].set_ylim(bottom=0)
fig.tight_layout()
pyplot.show()
class MichaelisMentenModel(murefi.BaseODEModel):
def __init__(self):
self.guesses = dict(S_0=5, P_0=0, v_max=0.1, K_S=1)
self.bounds = dict(
S_0=(1, 20),
P_0=(0, 10),
v_max=(0.0001, 5),
K_S=(0.01, 10),
)
super().__init__(
independent_keys=['S', 'P'],
parameter_names=["S_0", "P_0", "v_max", "K_S"],
)
def dydt(self, y, t, theta):
S, P = y
v_max, K_S = theta
dPdt = v_max * S / (K_S + S)
return [
-dPdt,
dPdt,
]
model = MichaelisMentenModel()
# generate ground truth data at 10 time points
theta_true = (8, 0, 0.16, 2.5)
rep_groundtruth = model.predict_replicate(
# S_0, P_0, v_max, K_S
parameters=theta_true,
template=murefi.Replicate.make_template(tmin=0, tmax=180, independent_keys='SP', rid='A01', N=10)
)
# use error model to make noisy observations of the ground truth
rep_observed = murefi.Replicate(rid='A01')
rep_observed['A570'] = murefi.Timeseries(
t=rep_groundtruth['P'].t,
y=observe_with_true_parameters(rep_groundtruth['P'].y),
independent_key='P',
dependent_key='A570'
)
# pack generated data into Dataset
dataset = murefi.Dataset()
dataset[rep_observed.rid] = rep_observed
fig, ax = pyplot.subplots(dpi=90)
ax.scatter(rep_observed['A570'].t, rep_observed['A570'].y, label='observations')
# use error model to project groundtruth onto dependent variable axis
ax.plot(rep_observed['A570'].t, cm_product.predict_dependent(rep_groundtruth["P"].y)[0], label='groundtruth')
ax.set(xlabel='time [s]', ylabel='$A_{570}$ [a.u.]')
ax.legend()
pyplot.show()
C:\Users\zufal\AppData\Local\Temp\ipykernel_9420\3436460410.py:33: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
fig.tight_layout()


Objective function and gradients
Like introduced in the basic example we need a ParameterMapping to specify how we want to apply the model to the dataset.
[3]:
df_mapping = pandas.DataFrame(columns='rid,S_0,P_0,v_max,K_S'.split(',')).set_index('rid')
# we'll fix S_0=8.0 and fit only the remaining parameters
df_mapping.loc['A01'] = (8.0, 'P_0', 'v_max', 'K_S')
# create the ParameterMapping object
pm = murefi.ParameterMapping(
df_mapping,
guesses=model.guesses, # fed as dict where the key is the name of the parameter
bounds=model.bounds # same as guesses
)
display(pm.as_dataframe())
display(pm)
| S_0 | P_0 | v_max | K_S | |
|---|---|---|---|---|
| rid | ||||
| A01 | 8.0 | P_0 | v_max | K_S |
ParameterMapping(1 replicates, 4 inputs, 3 free parameters)
The objective function is created in the same way:
[4]:
obj = murefi.objectives.for_dataset(
dataset,
model,
pm,
# no need for a substrate calibration model, because there's no data
calibration_models=[cm_product]
)
# The objective function can be evaluated at the initial guess to see if everything works as expected:
print(f'Negative log-likelihood at initial guess: {obj(pm.guesses)}')
Negative log-likelihood at initial guess: -11.26583475496314
With PyTensor installed, the objective can be called with TensorVariables to create a computation graph for the likelihood.
[5]:
K_S = pt.scalar("K_S")
P_0 = pt.scalar("P_0")
v_max = pt.scalar("v_max")
theta = [K_S, P_0, v_max]
L = obj([K_S, P_0, v_max])
# If `sunode` is installed we can auto-differentiate:
L_grad = pytensor.grad(L, wrt=theta)
# We can compile function to evaluate L and L_grad:
f_L = pytensor.function(inputs=theta, outputs=[L])
f_L_grad = pytensor.function(inputs=theta, outputs=L_grad)
parameters = pm.guesses
[6]:
print(f"""
Evaluations at pm.guesses:
obj : {obj(parameters)}
f_L : {f_L(*parameters)[0]}
f_L_grad : {f_L_grad(*parameters)}
""")
Evaluations at pm.guesses:
obj : -11.26583475496314
f_L : 11.265835029666126
f_L_grad : [array(-6.7969403), array(16.5343305), array(442.18509082)]
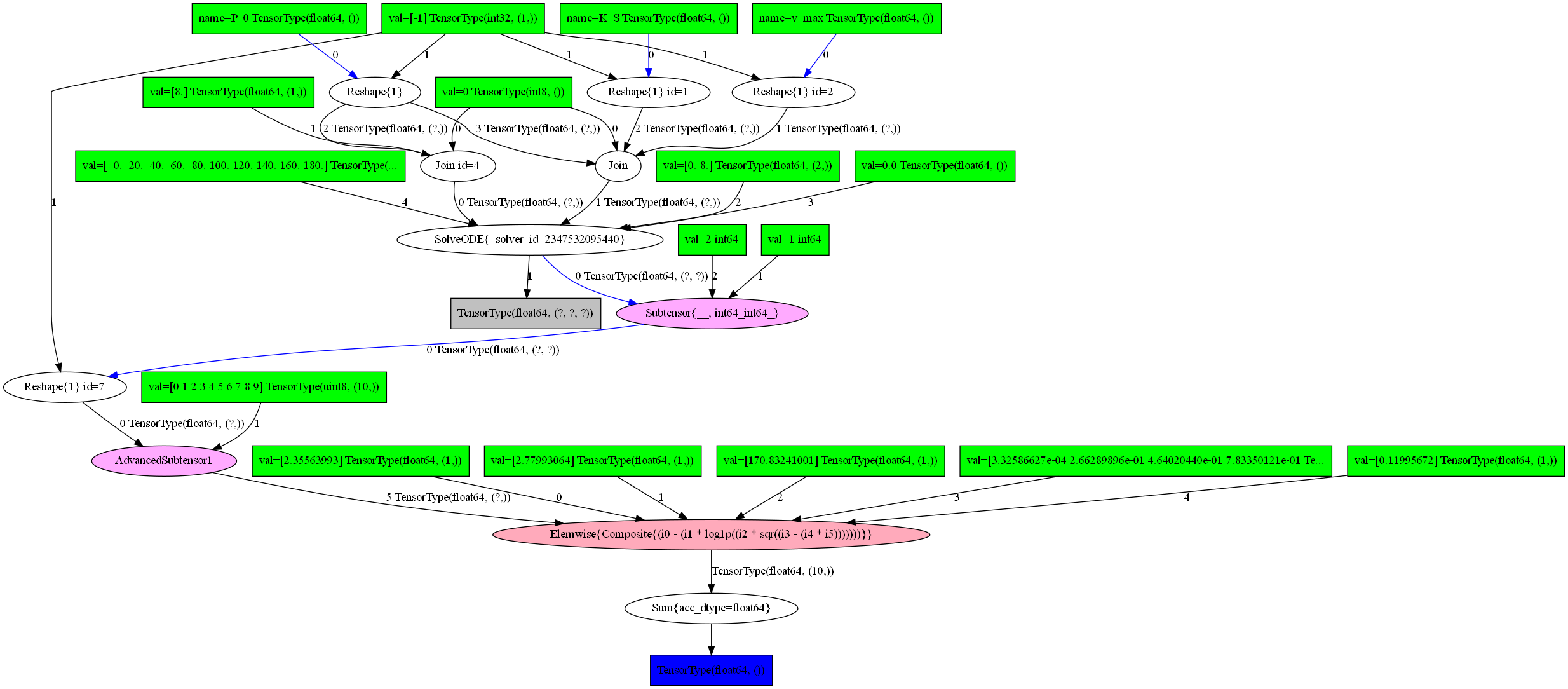
[7]:
# This cell needs graphviz and pydot:
# > mamba install -c conda-forge graphviz
# > pip install pydot
pytensor.printing.pydotprint(f_L, outfile="graph.png")
Image("graph.png")
The output file is available at graph.png
[7]:
Benchmarking
The compiled likelihood function with sunode should be faster than the odeint-based obj. At the very least the autograd gradient should be faster than a manual one.
[8]:
%%timeit
obj(parameters)
1.1 ms ± 16.4 µs per loop (mean ± std. dev. of 7 runs, 1,000 loops each)
[9]:
%%timeit
f_L(*parameters)
1.47 ms ± 34.8 µs per loop (mean ± std. dev. of 7 runs, 1,000 loops each)
[10]:
%%timeit
f_L_grad(*parameters)
1.45 ms ± 14.1 µs per loop (mean ± std. dev. of 7 runs, 1,000 loops each)
Building a Bayesian ODE model with murefi, calibr8, sunode and pymc
So far we have just seen how an ODE model built with murefi and calibr8 can be compiled with pytensor and auto-differentiated by pytensor and sunode. The above steps are good to know about for those who want to leverage fast likelihoods and gradients for other methods, but for building a PyMC model all of this happens behind the scenes.
Here we just have to come up with priors for our model parameters and can just press the inference button™.
[11]:
# first build the model
with pymc.Model() as pmodel:
t_theta = {
'S_0': 8,
'P_0': pymc.Exponential('P_0', lam=4),
'v_max': pymc.LogNormal('v_max', mu=numpy.log(3/25), sigma=1),
'K_S': pymc.LogNormal('K_S', mu=numpy.log(4), sigma=1)
}
# Just pass the priors into the objective.
# PyMC, sunode, murefi and calibr8 take care of the rest.
obj(t_theta)
pymc.model_to_graphviz(pmodel)
[11]:
[12]:
# then hit the inference button
with pmodel:
idata = pymc.sample(cores=1)
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [P_0, v_max, K_S]
Sampling 2 chains for 1_000 tune and 1_000 draw iterations (2_000 + 2_000 draws total) took 78 seconds.
[13]:
arviz.plot_trace(idata);
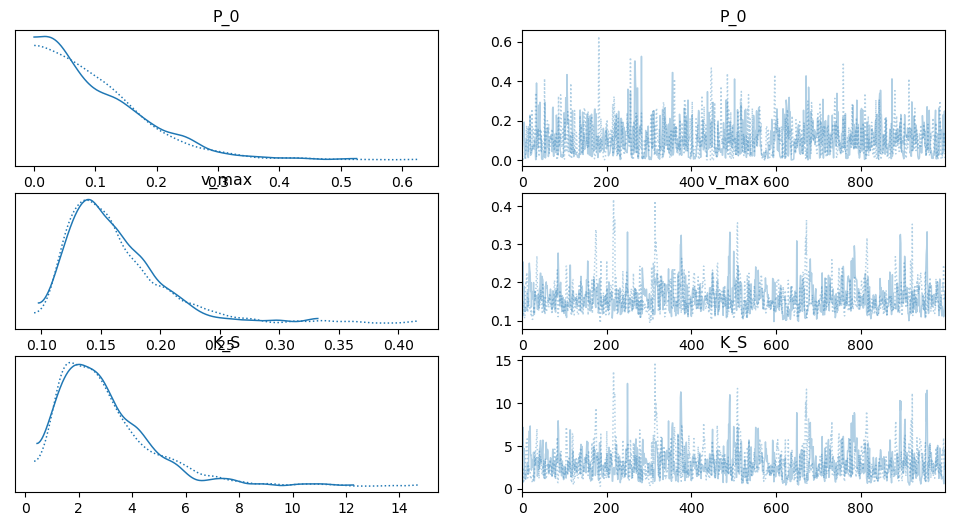
[14]:
dataset_pred = model.predict_dataset(
template=murefi.Dataset.make_template_like(dataset, independent_keys='SP'),
parameter_mapping=pm,
parameters={
k : idata.posterior[k].stack(sample=('chain', 'draw')).values
for k in pm.parameters.keys()
}
)
dataset_pred
[14]:
Dataset([('A01', Replicate(S[:200], P[:200]))])
[15]:
fig, ax = pyplot.subplots(dpi=90)
colors = dict(P='blue', S='orange')
pymc.gp.util.plot_gp_dist(
ax=ax,
x=dataset_pred['A01']['S'].t,
samples=dataset_pred['A01']['S'].y,
palette='Reds', plot_samples=False,
)
pymc.gp.util.plot_gp_dist(
ax=ax,
x=dataset_pred['A01']['P'].t,
samples=dataset_pred['A01']['P'].y,
palette='Blues', plot_samples=False,
)
# plot data, transformed via the error model into the independent unit
x_obs = rep_observed['A570'].t
y_obs = cm_product.predict_independent(rep_observed['A570'].y)
ax.scatter(x_obs, y_obs, label='observations',color=colors['P'])
# plot all timeseries of the groundtruth for comparison
for ykey, ts in reversed(rep_groundtruth.items()):
ax.plot(ts.t, ts.y, label=ykey + ' (true)', linestyle=':', color=colors[ykey])
ax.set_xlabel('time [s]')
ax.set_ylabel('P, S [mmol/L]')
ax.legend()
pyplot.show()
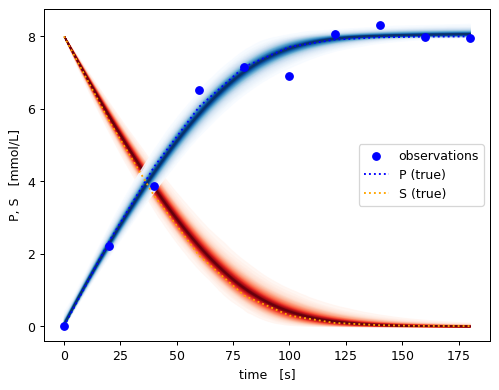
[16]:
%load_ext watermark
%watermark -n -u -v -iv -w
Last updated: Fri Dec 16 2022
Python implementation: CPython
Python version : 3.9.15
IPython version : 8.7.0
matplotlib: 3.6.2
pymc : 5.0.0
murefi : 5.2.0
pytensor : 2.8.10
pandas : 1.5.2
sunode : 0.4.0
calibr8 : 7.0.0
scipy : 1.9.3
arviz : 0.14.0
numpy : 1.23.5
Watermark: 2.3.1
[ ]: